scikit-image: image processing#
Author: Emmanuelle Gouillart
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
scikit-image is a Python package dedicated
to image processing, using NumPy arrays as image objects.
This chapter describes how to use scikit-image for various image
processing tasks, and how it relates to other scientific Python
modules such as NumPy and SciPy.
See also
For basic image manipulation, such as image cropping or simple filtering, a large number of simple operations can be realized with NumPy and SciPy only. See Image manipulation and processing using NumPy and SciPy.
Note that you should be familiar with the content of the previous chapter before reading the current one, as basic operations such as masking and labeling are a prerequisite.
Introduction and concepts#
Images are NumPy’s arrays np.ndarray
Pixels |
array values: |
Channels |
array dimensions |
Image encoding |
|
Filters |
functions ( |
# This example shows how to create a simple checkerboard.
check = np.zeros((8, 8))
check[::2, 1::2] = 1
check[1::2, ::2] = 1
plt.imshow(check, cmap='gray', interpolation='nearest');

scikit-image and the scientific Python ecosystem#
scikit-image is packaged in both pip and conda-based
Python installations, as well as in most Linux distributions. Other
Python packages for image processing & visualization that operate on
NumPy arrays include:
For N-dimensional arrays. Basic filtering, mathematical morphology, regions properties |
|
With a focus on high-speed implementations. |
|
A fast, interactive, multi-dimensional image viewer built in Qt. |
Some powerful C++ image processing libraries also have Python bindings:
A highly optimized computer vision library with a focus on real-time applications. |
|
The Insight ToolKit, especially useful for registration and working with 3D images. |
To varying degrees, these C++-based libraries tend to be less Pythonic and NumPy-friendly.
What is included in scikit-image#
Website: https://scikit-image.org/
Gallery of examples: https://scikit-image.org/docs/stable/auto_examples/
The library contains predominantly image processing algorithms, but also utility functions to ease data handling and processing. It contains the following submodules:
Color space conversion. |
|
Test images and example data. |
|
Drawing primitives (lines, text, etc.) that operate on NumPy arrays. |
|
Image intensity adjustment, e.g., histogram equalization, etc. |
|
Feature detection and extraction, e.g., texture analysis corners, etc. |
|
Sharpening, edge finding, rank filters, thresholding, etc. |
|
Graph-theoretic operations, e.g., shortest paths. |
|
Reading, saving, and displaying images and video. |
|
Measurement of image properties, e.g., region properties and contours. |
|
Metrics corresponding to images, e.g. distance metrics, similarity, etc. |
|
Morphological operations, e.g., opening or skeletonization. |
|
Restoration algorithms, e.g., deconvolution algorithms, denoising, etc. |
|
Partitioning an image into multiple regions. |
|
Geometric and other transforms, e.g., rotation or the Radon transform. |
|
Generic utilities. |
Importing#
We import scikit-image using the convention:
import skimage as ski
Most functionality lives in subpackages, e.g.:
image = ski.data.cat()
You can list all submodules with:
for m in dir(ski): print(m)
__version__
color
data
draw
exposure
feature
filters
future
graph
io
measure
metrics
morphology
registration
restoration
segmentation
transform
util
Most scikit-image functions take NumPy ndarrays as arguments
camera = ski.data.camera()
camera.dtype
dtype('uint8')
camera.shape
(512, 512)
filtered_camera = ski.filters.gaussian(camera, sigma=1)
type(filtered_camera)
numpy.ndarray
Example data#
To start off, we need example images to work with. The library ships with a few of these:
image = ski.data.cat()
image.shape
(300, 451, 3)
Input/output, data types and colorspaces#
I/O: skimage.io
Save an image to disk: skimage.io.imsave()
ski.io.imsave("cat.png", image)
Reading from files: skimage.io.imread()
cat = ski.io.imread("cat.png")
camera = ski.data.camera()
plt.figure(figsize=(4, 4))
plt.imshow(camera, cmap="gray", interpolation="nearest")
plt.axis("off")
plt.tight_layout()

This works with many data formats supported by the ImageIO library.
Loading also works with URLs:
logo = ski.io.imread('https://scikit-image.org/_static/img/logo.png')
Data types#
Image ndarrays can be represented either by integers (signed or unsigned) or floats.
Careful with overflows with integer data types
camera = ski.data.camera()
camera.dtype
camera_multiply = 3 * camera

Different integer sizes are possible: 8-, 16- or 32-bytes, signed or unsigned.
Warning: An important (if questionable) skimage convention: float
images are supposed to lie in [-1, 1] (in order to have comparable contrast
for all float images):
camera_float = ski.util.img_as_float(camera)
camera.max(), camera_float.max()
(np.uint8(255), np.float64(1.0))
Some image processing routines need to work with float arrays, and may hence output an array with a different type and the data range from the input array
camera_sobel = ski.filters.sobel(camera)
camera_sobel.max()
np.float64(0.6447887988758096)
Utility functions are provided in skimage to convert both the
dtype and the data range, following skimage’s conventions:
util.img_as_float, util.img_as_ubyte, etc.
See the user guide for more details.
Colorspaces#
Color images are of shape (N, M, 3) or (N, M, 4) (when an alpha channel encodes transparency)
face = sp.datasets.face()
face.shape
(768, 1024, 3)
Routines converting between different colorspaces (RGB, HSV, LAB etc.)
are available in skimage.color : color.rgb2hsv, color.lab2rgb,
etc. Check the docstring for the expected dtype (and data range) of input
images.
3D images
Most functions of skimage can take 3D images as input arguments.
Check the docstring to know if a function can be used on 3D images
(for example MRI or CT images).
Exercise 68
Open a color image on your disk as a NumPy array.
Find a skimage function computing the histogram of an image and plot the histogram of each color channel
Convert the image to grayscale and plot its histogram.
Image preprocessing / enhancement#
Goals: denoising, feature (edges) extraction, …
Local filters#
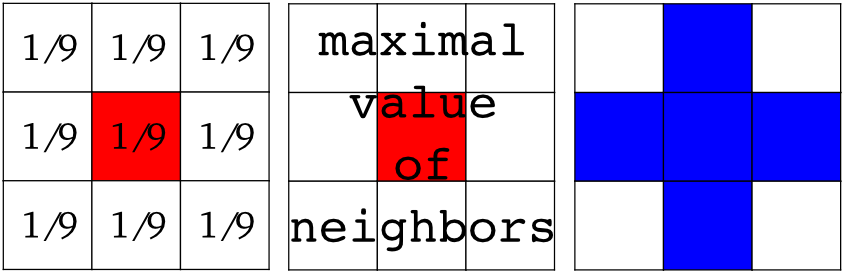
Local filters replace the value of pixels by a function of the values of neighboring pixels. The function can be linear or non-linear.
Neighbourhood: square (choose size), disk, or more complicated structuring element.

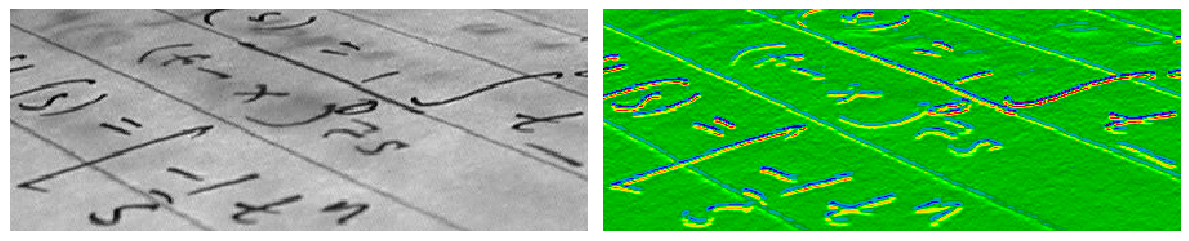
Example : horizontal Sobel filter
text = ski.data.text()
hsobel_text = ski.filters.sobel_h(text)
Uses the following linear kernel for computing horizontal gradients:
1 2 1
0 0 0
-1 -2 -1

Non-local filters#
Non-local filters use a large region of the image (or all the image) to transform the value of one pixel:
camera = ski.data.camera()
camera_equalized = ski.exposure.equalize_hist(camera)
Enhances contrast in large almost uniform regions.

Mathematical morphology#
See wikipedia for an introduction on mathematical morphology.
Probe an image with a simple shape (a structuring element), and modify this image according to how the shape locally fits or misses the image.
Default structuring element: 4-connectivity of a pixel
# Import structuring elements to make them more easily accessible
from skimage.morphology import disk, diamond
diamond(1)
array([[0, 1, 0],
[1, 1, 1],
[0, 1, 0]], dtype=uint8)
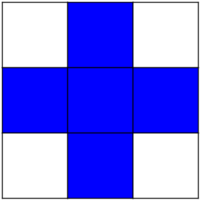
Erosion = minimum filter. Replace the value of a pixel by the minimal value covered by the structuring element.:
a = np.zeros((7,7), dtype=np.uint8)
a[1:6, 2:5] = 1
a
array([[0, 0, 0, 0, 0, 0, 0],
[0, 0, 1, 1, 1, 0, 0],
[0, 0, 1, 1, 1, 0, 0],
[0, 0, 1, 1, 1, 0, 0],
[0, 0, 1, 1, 1, 0, 0],
[0, 0, 1, 1, 1, 0, 0],
[0, 0, 0, 0, 0, 0, 0]], dtype=uint8)
ski.morphology.binary_erosion(a, diamond(1)).astype(np.uint8)
/tmp/ipykernel_7509/4035965298.py:1: FutureWarning: `binary_erosion` is deprecated since version 0.26 and will be removed in version 0.28. Use `skimage.morphology.erosion` instead. Note the pixel shift by 1 for even-sized footprints (see docstring notes).
ski.morphology.binary_erosion(a, diamond(1)).astype(np.uint8)
array([[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 1, 0, 0, 0],
[0, 0, 0, 1, 0, 0, 0],
[0, 0, 0, 1, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0]], dtype=uint8)
#Erosion removes objects smaller than the structure
ski.morphology.binary_erosion(a, diamond(2)).astype(np.uint8)
/tmp/ipykernel_7509/3101513642.py:2: FutureWarning: `binary_erosion` is deprecated since version 0.26 and will be removed in version 0.28. Use `skimage.morphology.erosion` instead. Note the pixel shift by 1 for even-sized footprints (see docstring notes).
ski.morphology.binary_erosion(a, diamond(2)).astype(np.uint8)
array([[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 0]], dtype=uint8)
Dilation: maximum filter:
a = np.zeros((5, 5))
a[2, 2] = 1
a
array([[0., 0., 0., 0., 0.],
[0., 0., 0., 0., 0.],
[0., 0., 1., 0., 0.],
[0., 0., 0., 0., 0.],
[0., 0., 0., 0., 0.]])
ski.morphology.binary_dilation(a, diamond(1)).astype(np.uint8)
/tmp/ipykernel_7509/3795506340.py:1: FutureWarning: `binary_dilation` is deprecated since version 0.26 and will be removed in version 0.28. Use `skimage.morphology.dilation` instead. Note the lack of mirroring for non-symmetric footprints (see docstring notes).
ski.morphology.binary_dilation(a, diamond(1)).astype(np.uint8)
array([[0, 0, 0, 0, 0],
[0, 0, 1, 0, 0],
[0, 1, 1, 1, 0],
[0, 0, 1, 0, 0],
[0, 0, 0, 0, 0]], dtype=uint8)
Opening: erosion + dilation:
a = np.zeros((5,5), dtype=int)
a[1:4, 1:4] = 1; a[4, 4] = 1
a
array([[0, 0, 0, 0, 0],
[0, 1, 1, 1, 0],
[0, 1, 1, 1, 0],
[0, 1, 1, 1, 0],
[0, 0, 0, 0, 1]])
ski.morphology.binary_opening(a, diamond(1)).astype(np.uint8)
/tmp/ipykernel_7509/2186513000.py:1: FutureWarning: `binary_opening` is deprecated since version 0.26 and will be removed in version 0.28. Use `skimage.morphology.opening` instead.
ski.morphology.binary_opening(a, diamond(1)).astype(np.uint8)
array([[0, 0, 0, 0, 0],
[0, 0, 1, 0, 0],
[0, 1, 1, 1, 0],
[0, 0, 1, 0, 0],
[0, 0, 0, 0, 0]], dtype=uint8)
Opening removes small objects and smoothes corners.
Grayscale mathematical morphology
Mathematical morphology operations are also available for (non-binary) grayscale images (int or float type). Erosion and dilation correspond to minimum (resp. maximum) filters.
Higher-level mathematical morphology are available: tophat, skeletonization, etc.
See also
Basic mathematical morphology is also implemented in
scipy.ndimage.morphology. The scipy.ndimage implementation
works on arbitrary-dimensional arrays.
Example of filters comparison: image denoising#
coins = ski.data.coins()
coins_zoom = coins[10:80, 300:370]
median_coins = ski.filters.median(
coins_zoom, disk(1)
)
tv_coins = ski.restoration.denoise_tv_chambolle(
coins_zoom, weight=0.1
)
gaussian_filter_coins = ski.filters.gaussian(coins, sigma=2)
med_filter_coins = ski.filters.median(coins, np.ones((3, 3)))
tv_filter_coins = ski.restoration.denoise_tv_chambolle(coins, weight=0.1)
Text(0.5, 1.0, 'TV filter')

Image segmentation#
Image segmentation is the attribution of different labels to different regions of the image, for example in order to extract the pixels of an object of interest.
Binary segmentation: foreground + background#
Histogram-based method: Otsu thresholding#
Note
The Otsu method is a simple heuristic to find a threshold to separate the foreground from the background.
camera = ski.data.camera()
val = ski.filters.threshold_otsu(camera)
mask = camera < val
# The histogram from which Otsu calculated the threshold.
hist, bins_center = ski.exposure.histogram(camera)
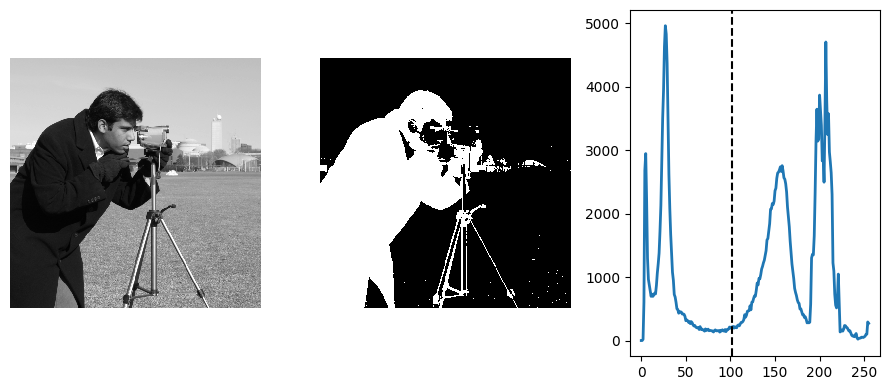

Labeling connected components of a discrete image#
Note
Once you have separated foreground objects, it is use to separate them from each other. For this, we can assign a different integer labels to each one.
Synthetic data:
n = 20
l = 256
im = np.zeros((l, l))
rng = np.random.default_rng()
points = l * rng.random((2, n ** 2))
im[(points[0]).astype(int), (points[1]).astype(int)] = 1
im = ski.filters.gaussian(im, sigma=l / (4. * n))
blobs = im > im.mean()
Label all connected components:
all_labels = ski.measure.label(blobs)
Label only foreground connected components:
blobs_labels = ski.measure.label(blobs, background=0)
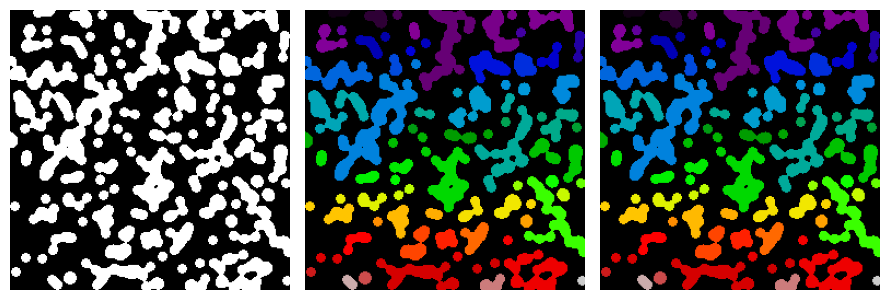

See also
scipy.ndimage.find_objects() is useful to return slices on
object in an image.
Marker based methods#
If you have markers inside a set of regions, you can use these to segment the regions.
Watershed segmentation#
The Watershed (skimage.segmentation.watershed()) is a region-growing
approach that fills “basins” in the image
# Generate an initial image with two overlapping circles
x, y = np.indices((80, 80))
x1, y1, x2, y2 = 28, 28, 44, 52
r1, r2 = 16, 20
mask_circle1 = (x - x1) ** 2 + (y - y1) ** 2 < r1 ** 2
mask_circle2 = (x - x2) ** 2 + (y - y2) ** 2 < r2 ** 2
image = np.logical_or(mask_circle1, mask_circle2)
# Now we want to separate the two objects in image
# Generate the markers as local maxima of the distance
# to the background
# Use scipy.ndimage.distance_transform_edt
distance = sp.ndimage.distance_transform_edt(image)
peak_idx = ski.feature.peak_local_max(
distance, footprint=np.ones((3, 3)), labels=image
)
peak_mask = np.zeros_like(distance, dtype=bool)
peak_mask[tuple(peak_idx.T)] = True
markers = ski.morphology.label(peak_mask)
labels_ws = ski.segmentation.watershed(
-distance, markers, mask=image
)
Random walker segmentation#
The random walker algorithm (skimage.segmentation.random_walker())
is similar to the Watershed, but with a more “probabilistic” approach. It
is based on the idea of the diffusion of labels in the image:
# Transform markers image so that 0-valued pixels are to
# be labelled, and -1-valued pixels represent background
markers[~image] = -1
labels_rw = ski.segmentation.random_walker(image, markers)
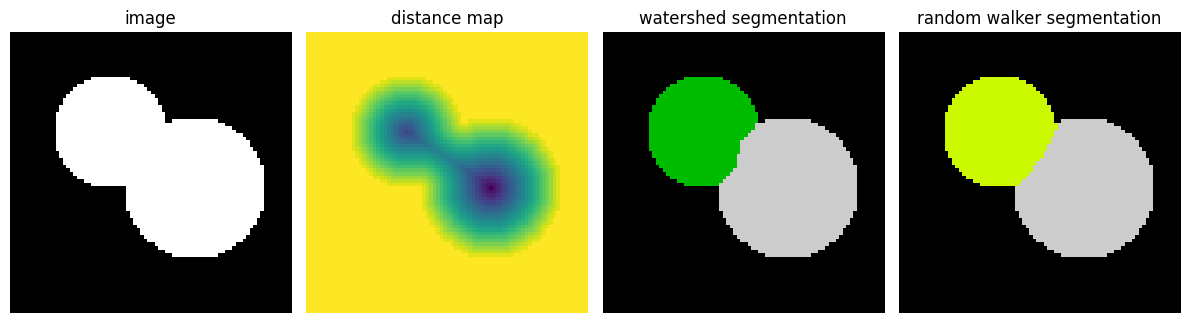
Postprocessing label images
skimage provides several utility functions that can be used on
label images (ie images where different discrete values identify
different regions). Functions names are often self-explaining:
skimage.segmentation.clear_border(),
skimage.segmentation.relabel_from_one(),
skimage.morphology.remove_small_objects(), etc.
Exercise 69
Load the
coinsimage from thedatasubmodule.Separate the coins from the background by testing several segmentation methods: Otsu thresholding, adaptive thresholding, and watershed or random walker segmentation.
If necessary, use a postprocessing function to improve the coins / background segmentation.
Measuring regions’ properties#
Example: compute the size and perimeter of the two segmented regions:
properties = ski.measure.regionprops(labels_rw)
[float(prop.area) for prop in properties]
[770.0, 1168.0]
[prop.perimeter for prop in properties]
[np.float64(100.91168824543142), np.float64(126.81118318204308)]
See also
for some properties, functions are available as well in
scipy.ndimage.measurements with a different API (a list is
returned).
Exercise 70
Use the binary image of the coins and background from the previous exercise.
Compute an image of labels for the different coins.
Compute the size and eccentricity of all coins.
Data visualization and interaction#
Meaningful visualizations are useful when testing a given processing pipeline.
Some image processing operations:
coins = ski.data.coins()
mask = coins > ski.filters.threshold_otsu(coins)
clean_border = ski.segmentation.clear_border(mask)
Visualize binary result:
plt.figure()
plt.imshow(clean_border, cmap='gray')
<matplotlib.image.AxesImage at 0x7f2030d21370>
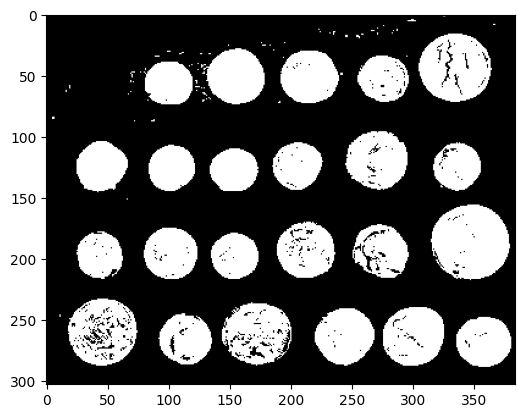

Visualize contour
plt.figure()
plt.imshow(coins, cmap='gray')
plt.contour(clean_border, [0.5])
<matplotlib.contour.QuadContourSet at 0x7f2030f959d0>
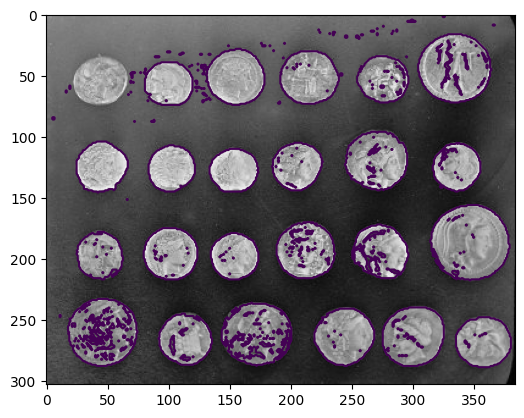

Use skimage dedicated utility function:
coins_edges = ski.segmentation.mark_boundaries(
coins, clean_border.astype(int)
)
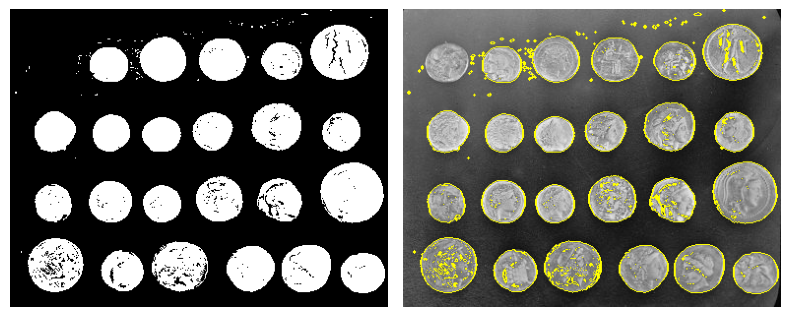
Feature extraction for computer vision#
Geometric or textural descriptor can be extracted from images in order to
classify parts of the image (e.g. sky vs. buildings)
match parts of different images (e.g. for object detection)
and many other applications of Computer Vision
Example: detecting corners using Harris detector
tform = ski.transform.AffineTransform(
scale=(1.3, 1.1), rotation=1, shear=0.7,
translation=(210, 50)
)
image = ski.transform.warp(
ski.data.checkerboard(), tform.inverse, output_shape=(350, 350)
)
coords = ski.feature.corner_peaks(
ski.feature.corner_harris(image), min_distance=5
)
coords_subpix = ski.feature.corner_subpix(
image, coords, window_size=13
)
(np.float64(-23.504322001100714),
np.float64(349.5),
np.float64(349.5),
np.float64(-0.5))
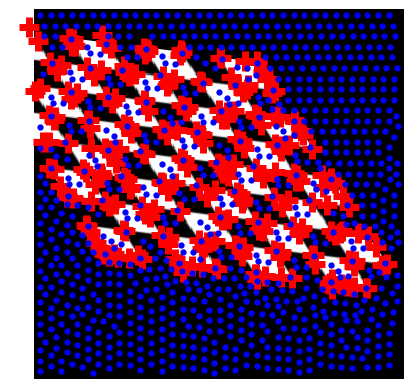
(this example is taken from the plot_corner example in scikit-image)
Points of interest such as corners can then be used to match objects in different images, as described in the plot_matching example of scikit-image.

